Research
One third of the human population carries the invasive pathogen Staphylococcus aureus within the nasal cavity. This colonization increases the risk of infection and antibiotic resistant lineages (especially MRSA) put urgency on the development of new strategies to prevent and treat infections. The Heilbronner lab addresses this problem in three distinct research topics.
1) Virulence: We investigate the molecular mechanisms of S. aureus pathogenicity during invasive disease with a special focus on trace metal acquisition (read more)
2) Interaction with the Microbiota: We investigate if bacteria of the nasal microbiome can provide resilience against S. aureus colonization (read more)
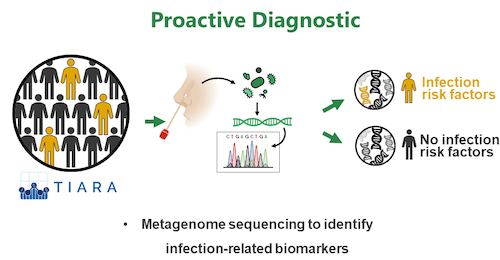
3) Proactive Diagnostic: Using clinical cohorts, we investigate if metagenome sequencing of nasal specimen can identify microbiome signatures (biomarkers) predicting the risk of a patient to be infected by S. aureus (read more)
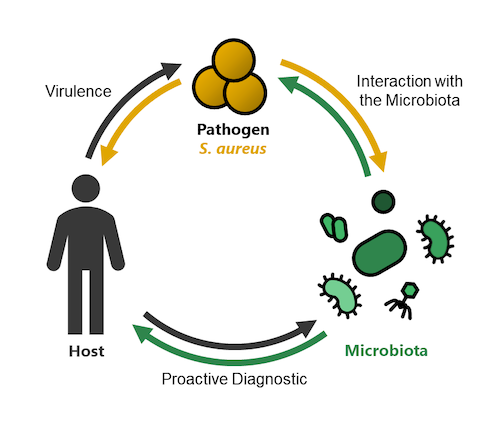
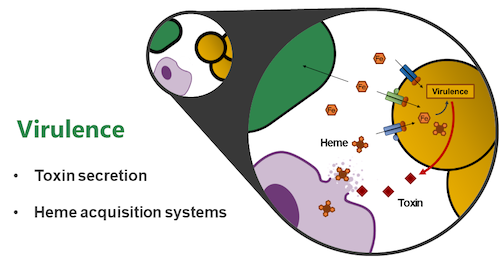
Research Topic 1: Molecular Mechanisms of Staphylococcal Virulence
In order to cause infection, pathogens need to overcome humoral and cellular immune defenses of the host. Additionally, pathogens need to acquire host-derived nutrients and adhere to extracellular matrix molecules.
Our major focus lies on the molecular mechanisms of staphylococcal trace metal acquisition during infection. Iron is essential for bacterial proliferation and the active limitation of iron within the human body is an important immune effector mechanism. However, within humans, iron-containing heme is used for oxygen transport and pathogens have developed complex systems to acquire and degrade heme. Staphylococci secrete cytolytic toxins to release hemoproteins from host cells. In a second step, complex systems employing cell wall-anchored receptors, and membrane ABC-transporters allow coordinated transport of heme across the cell envelope. We study the molecular architecture of these systems on the cellular level using state of the art genetic, biochemical, and in vivo approaches.
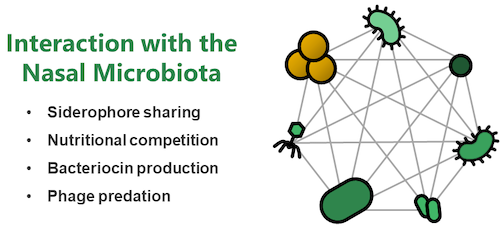
Research Topic 2: Molecular Interactions with the Nasal Microbiome
Nasal colonization of S. aureus represents a major reservoir for infection. However, it is incompletely understood why some human individuals carry S. aureus while others show resilience towards colonization. Within the nasal cavity S. aureus needs to interact with other bacteria of the nasal microbiome and bacterial interactions can be of collaborative or competitive nature. We use extensive strain collections of nasal bacterial isolates and study their interaction with S. aureus. We use high-throughput approaches to identify networks of reciprocal collaboration and support and investigate the underlying molecular mechanisms. Of major focus are interactions that are based on shared nutritional resources (mucins, iron-binding siderophores etc.), bacteriocins (antibacterial molecules) and bacteriophages (viruses infecting bacteria). Hypothesis deriving from our in vitro experiments are tested in mice harboring a humanized nasal microbiome. Our overarching aim is to develop nasal probiotics to displace the pathogen from the nasal cavity of humans.
Research Topic 3: Proactive Diagnostics
It is known that S. aureus nasal colonization represents a risk factor for infection with the endogenous strain. However, direct means to translate this knowledge into clinical applications is currently lacking. In frame of the German Centre For Infection Research (DZIF) we use clinical cohorts and full metagenome sequencing of nasal specimen to identify microbiome signatures (e.g. pathogen abundance, presence/absence of specific commensals) that correlate with the development of infection. Such biomarkers are of high relevance as they can encourage proactive treatment of certain patients before infection occurs.